| CD200 |
|---|
 |
|
| Structures disponibles |
|---|
| PDB | Recherche d'orthologue: PDBe RCSB |
|---|
|
|
| Identifiants |
|---|
| Aliases | CD200 |
|---|
| IDs externes | OMIM: 155970 MGI: 1196990 HomoloGene: 4344 GeneCards: CD200 |
|---|
| Position du gène (Homme) |
|---|
 | | Chr. | Chromosome 3 humain[1] |
|---|
| | Locus | 3q13.2 | Début | 112,332,347 bp[1] |
|---|
| Fin | 112,362,812 bp[1] |
|---|
|
| Position du gène (Souris) |
|---|
 | | Chr. | Chromosome 16 (souris)[2] |
|---|
| | Locus | 16 B5|16 29.53 cM | Début | 45,202,498 bp[2] |
|---|
| Fin | 45,229,416 bp[2] |
|---|
|
| Expression génétique |
|---|
| Bgee | | Humain | Souris (orthologue) |
|---|
| Fortement exprimé dans | - frontal pole
- éminence ganglionnaire
- Brodmann area 10
- gyrus frontal moyen
- hypothalamus
- cortex orbitofrontal
- Brodmann area 9
- parietal pleura
- Brodmann area 46
- Brodmann area 23
|
| | Fortement exprimé dans | - aorte ascendante
- tunica media of zone of aorta
- Aire tegmentale ventrale
- dorsomedial hypothalamic nucleus
- ventromedial nucleus
- noyau paraventriculaire de l'hypothalamus
- Noyau suprachiasmatique
- corps du fémur
- noyau arqué
- valve aortique
|
| | Plus de données d'expression de référence |
|
|---|
| BioGPS | 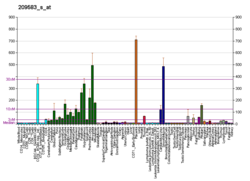
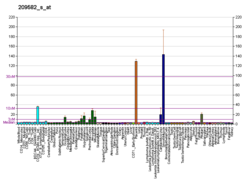 | | Plus de données d'expression de référence |
|
|---|
|
| Gene Ontology |
|---|
| Fonction moléculaire | - signaling receptor binding
- liaison protéique
- protein homodimerization activity
- cell adhesion molecule binding
- protein binding involved in heterotypic cell-cell adhesion
- glycosylated region protein binding
| | Composant cellulaire | - integral component of membrane
- integral component of plasma membrane
- membrane
- membrane plasmique
- surface cellulaire
- axone
- neuron projection
- neuronal cell body
- cell body
| | Processus biologique | - heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules
- cell recognition
- homophilic cell adhesion via plasma membrane adhesion molecules
- negative regulation of macrophage activation
- regulation of immune response
- negative regulation of leukocyte activation
- negative regulation of cell population proliferation
- negative regulation of NF-kappaB transcription factor activity
- positive regulation of CREB transcription factor activity
- heterotypic cell-cell adhesion
- positive regulation of transforming growth factor beta production
- cell-cell adhesion
- positive regulation of arginase activity
- positive regulation of protein-glutamine gamma-glutamyltransferase activity
- regulation of neuroinflammatory response
- negative regulation of neuroinflammatory response
- negative regulation of neuron death
- negative regulation of matrix metallopeptidase secretion
- negative regulation of macrophage migration
- negative regulation of T cell migration
| | Sources:Amigo / QuickGO |
|
| Orthologues |
|---|
| Espèces | Homme | Souris |
|---|
| Entrez | | |
|---|
| Ensembl | | |
|---|
| UniProt | | |
|---|
| RefSeq (mRNA) | NM_001004196
NM_001004197
NM_005944
NM_001318826
NM_001318828
|
|---|
NM_001318830
NM_001365851
NM_001365852
NM_001365853
NM_001365854
NM_001365855 |
| |
|---|
| RefSeq (protéine) | NP_001004196
NP_001305755
NP_001305757
NP_001305759
NP_005935
|
|---|
NP_001352780
NP_001352781
NP_001352782
NP_001352783
NP_001352784 |
| |
|---|
| Localisation (UCSC) | Chr 3: 112.33 – 112.36 Mb | Chr 16: 45.2 – 45.23 Mb |
|---|
| Publication PubMed | [3] | [4] |
|---|
|
| Wikidata |
| Voir/Editer Humain | Voir/Editer Souris |
|


 Portail de la médecine
Portail de la médecine 
















